An innovative study on DNAThe hidden structures of the brain could open up new approaches to treating and diagnosing diseases, including cancer.
Researchers at the Garvan Institute have published the first comprehensive map of over 50,000 i-motifs in the human genome – structures that differ from the classic double helix and may play a critical role in gene regulation and disease. These findings highlight the potential of i-motifs in developing new therapies, particularly in targeting genes associated with cancer.
Unlocking the secrets of DNA i-motifs
DNA is known for its double helix shape, but researchers at the Garvan Institute of Medical Research have discovered that the human genome also contains over 50,000 unusual “knot-like” DNA structures called i-motifs.
Published today (29 August) in The EMBO Journal is the first comprehensive map of these unique DNA structures, shedding light on their potential role in gene regulation related to disease.
In a groundbreaking study in 2018, Garvan scientists were the first to visualize i-motifs directly in living human cells using a new antibody tool they developed to detect and bind i-motifs. The current research builds on these findings and uses this antibody to identify i-motif locations across the genome.
“In this study, we have mapped more than 50,000 i-motif sites in the human genome, occurring in all three cell types we studied,” says lead author Professor Daniel Christ, Head of the Antibody Therapeutics Lab and Director of the Centre for Targeted Therapy at Garvan. “This is a remarkably high number for a DNA structure whose existence in cells was once considered controversial. Our results confirm that i-motifs are not just laboratory curiosities, but are widespread – and likely play a key role in genome function.”
Key roles of i-motifs in gene regulation
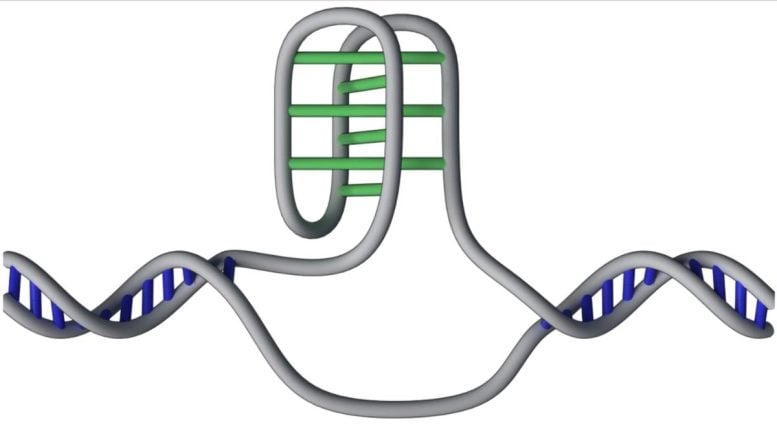
I-motifs are DNA structures that deviate from the typical double helix shape. They are formed when stretches of cytosine letters on the same DNA strand pair together, creating a four-stranded, twisted structure that protrudes from the double helix.
Animation of the knot-like i-motif structure of DNA, which the team mapped to 50,000 locations in the human genome, focused on key functional areas, including regions that control gene activity. Image credit: Cristian David Pena Martinez / Garvan Institute
The researchers found that i-motifs are not randomly scattered but are concentrated in important functional areas of the genome, including regions that control gene activity.
“We found that i-motifs are associated with genes that are highly active at certain times in the cell cycle, suggesting that they play a dynamic role in regulating gene activity,” says Cristian David Peña Martinez, research associate in the Antibody Therapeutics Lab and lead author of the study.
“We also found that i-motifs form in the promoter region of oncogenes, for example in the MyC oncogene that encodes one of the most notorious ‘undruggable’ targets of cancer. This presents an exciting opportunity to target disease-associated genes via the i-motif structure,” he says.

Therapeutic potential of i-motifs
“The widespread occurrence of i-motifs near these ‘holy grail’ sequences involved in difficult-to-treat cancers opens up new opportunities for new diagnostic and therapeutic approaches. It may be possible to develop drugs that target i-motifs to influence gene expression, which could expand current treatment options,” says Associate Professor Sarah Kummerfeld, Chief Scientific Officer at Garvan and co-author of the study.
Professor Christ adds that mapping i-motifs was only possible thanks to Garvan’s world-leading expertise in antibody development and genomics. “This study is an example of how basic research and technological innovation can come together to enable groundbreaking discoveries,” he says.
Reference: “Human genomic DNA is widely interspersed with i-motif structures” August 29, 2024, The EMBO Journal.
DOI: 10.1038/s44318-024-00210-5
Professor Daniel Christ is an Associate Professor at St Vincent’s Clinical School, Faculty of Medicine and Health, UNSW Sydney. Associate Professor Sarah Kummerfeld is an Associate Professor at St Vincent’s Clinical School, Faculty of Medicine and Health, UNSW Sydney.
This research was supported by funding from the National Health and Medical Research Council.

